|
This style of copying is useful when interacting with programs like Sequencher (TM). That is, whereas the standard Copy would place into the clipboard selected pieces of the matrix in tab-delimited text format (e.g., if the sequence AATCA is selected, "A-tab-A-tab-T-tab-C-tab-A" would be copied), this modified Copy Sequence command does not include tabs (thus, "AATCA" would be copied). The Display menu contains other options such as a Bird's eye view which makes the cells narrow to show more of the sequences.Ĭopy Sequence (at bottom of Edit menu) copies the selected cells of the matrix into the computer's clipboard as a sequence. Cells can be colored according to the state at the site (Color Cells submenu, Character State) or according to a value like the GC bias (Color Cells submenu,Cell Value can request this coloring to use a moving window). The view of the matrix can be adjusted in various ways in the Display menu. You can also apply the other editing tools described for character matrices. Other options may appear see the page on characters for standard choices in this submenu. Trim Terminal Gap Characters - deletes characters at edges of matrix that are gaps-only.Nucleotide complement (DNA matrix only) - enters the complementary sequence into the selected cells.Thus, a Y will be converted to a C with 50% probability, and to a T with 50%. Arbitrarily Resolve State Ambiguity - Any cell that has partial uncertainty in the state (e.g., "Y", which is C or T) will be resolved into one of its states, chosen randomly.Convert to RY - Converts A and G to R, and C and T, in a nucleotide sequence.For example, for a nucleotide sequence, a cell that has both C and T will be converted into "C or T", i.e., Y. Convert Polymorphisms to Uncertainties - Converts all polymorphisms in the selection to uncertainty. 
For example, for a nucleotide sequence, a Y (C or T) will be converted into "C andT". 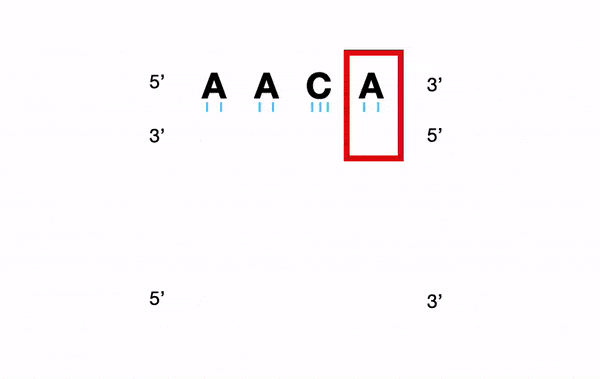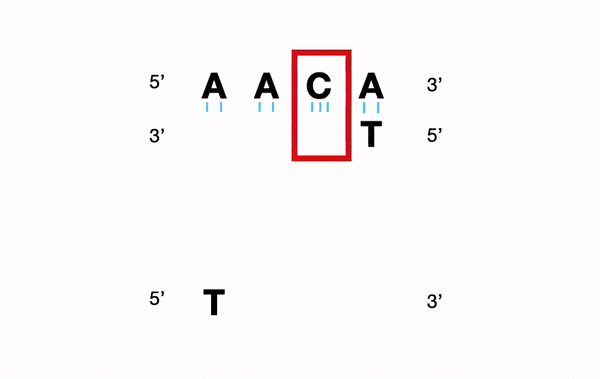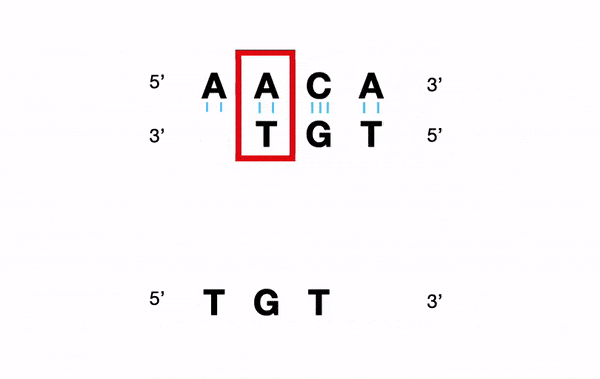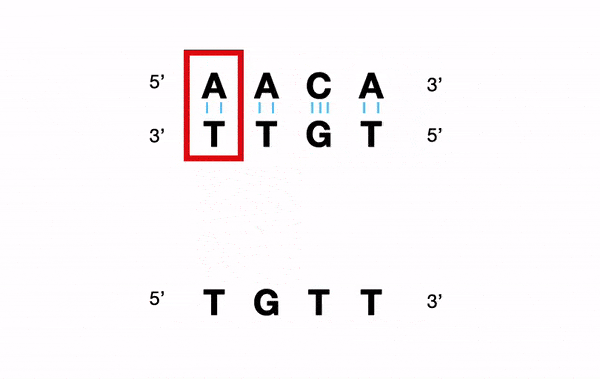
Convert Uncertainties to Polymorphisms - Converts all uncertainties in the selection to polymorphisms.This feature requires that codon positions be designated. The amount each sequence will be shifted will vary from sequence to sequence. Shift To Minimize Stop Codons - Shifts each sequence 0, 1, or 2 bases so as to minimize the number of stop codons.Shift Other to Match - Shifts other sequences to match a region in a selected sequences described in the manual for the Align package.Remove Gaps-Only Characters - Removes from the matrix all characters that consist of nothing but gaps.Collapse Gaps - collapses gaps in the selected block by pushing all sites to the left, to to yield unaligned sequences.Reverse Complement (DNA matrix only) - reverses the order of contiguously selected blocks of sequence and complements the sequence.These are available under the Alter/Transform submenu of the Matrix menu (some of these may be under "Other Choices"): The following can be applied to all or the selected portions of a molecular sequence matrix in the Character Matrix Editor. The Character Matrix Editor can be used to edit a molecular sequence matrix. It can also be exported to some of these formats. 
Molecular data can be imported from files of NBRF, FASTA, GenBank/GenPept, PHYLIP, CLUSTAL, and simple table format.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
